Professional, Customizable Oryza sativa Transformation Services
Lifeasible is a recognized leader in plant biotechnology, offering a comprehensive and high-efficiency platform for rice transformation service. Oryza sativa (rice) serves as the primary staple food for more than half of the global population and remains the premier model organism for monocot functional genomics and molecular breeding. Our services are designed to overcome the technical barriers associated with rice regeneration, providing academic researchers and the AgBio industry with a streamlined path from gene concept to stable transgenic events.
Leveraging our deep expertise in plant genetic engineering, we provide end-to-end support for a variety of projects, including nutritional biofortification, yield enhancement, and the development of biotic and abiotic stress resistance.
TARGET GENOTYPES
Japonica & Indica
Nipponbare, Minghui 63, etc.
TYPICAL YIELD
10-30
Independent T0 Positive Rice Events
EDITING EFFICIENCY
Up to 80%
High-efficiency CRISPR/Cas9 gene editing
LEAD TIME
4-6 Months
From vector receipt to T1 seeds
Standard Package
Efficiency Focused
Premium Package
Full-Service Custody
Stable transformation is the bedrock of modern rice improvement, enabling the permanent integration and inheritance of novel genetic traits. At Lifeasible, we have optimized the Agrobacterium-mediated transformation process to ensure high-frequency T-DNA integration with a high proportion of single-copy events.
While Agrobacterium is our primary method due to its clean integration patterns, we also offer other transformation methods for specialized applications, such as plant chloroplast genome editing or when working with high-molecular-weight DNA constructs that exceed T-DNA carrying capacities.
![]()
Explant Selection
Use of high-quality mature seeds or immature embryos to induce embryogenic calli.
![]()
Infection & Co-cultivation
Precise timing of Agrobacterium inoculation with specialized acetosyringone-enhanced media to boost vir gene induction.
![]()
Stringent Selection
Multi-step antibiotic selection (e.g., Hygromycin, G418) to eliminate non-transgenic tissues while maintaining callus vitality.
![]()
Regeneration
Optimized hormonal ratios to trigger shoot and root development, minimizing somaclonal variation.
![]()
Acclimatization
Controlled greenhouse hardening to ensure a high survival rate of T0 plantlets.
For projects requiring rapid data turnaround, Lifeasible provides high-throughput transient expression systems that bypass the lengthy regeneration phase. These assays allow for the functional validation of gene constructs, promoter strength analysis, or subcellular localization in days rather than months, providing a critical fast-track for preliminary research before committing to stable transformation.
![]()
Vector Design & Preparation
Selection of optimized transient expression vectors (e.g., pHBT, pUC) and high-purity plasmid extraction.
![]()
Target Material Isolation
Preparation of high-viability rice protoplasts from etiolated seedlings or healthy leaf tissues for viral/bombardment assays.
![]()
DNA Delivery
Application of PEG-mediated, virus-mediated, or biolistic delivery methods.
![]()
Incubation & Analysis
Controlled cultivation followed by fluorescence imaging, qPCR, or Western Blotting.
Lifeasible employs a diverse and optimized toolkit to overcome the challenges associated with monocot genetic engineering. We offer a selection of transformation methodologies to ensure successful DNA delivery into Oryza sativa tissues, catering to both stable integration and transient functional analysis requirements.
This is our primary method for generating stable transgenic rice lines. We utilize optimized Agrobacterium tumefaciens strains (e.g., EHA105, AGL1, or GV3101) and virulence-enhancing compounds such as acetosyringone to infect mature seed-derived embryogenic callus. This method is preferred for its ability to produce transgenic plants with low copy numbers and stable inheritance of the target gene.
We utilize plant viral vectors to facilitate rapid gene function analysis in rice. This method is particularly powerful for Virus-Induced Gene Silencing (VIGS) and virus-induced gene spreading, allowing researchers to quickly assess loss-of-function or gain-of-function phenotypes in rice seedlings or specific tissues without the extensive timeline required for generating stable mutants.
PEG-mediated transformation is a high-efficiency chemical method used to induce direct DNA uptake. At Lifeasible, this technique is predominantly applied to rice protoplasts isolated from etiolated seedlings or suspension cells. It serves as an ideal platform for high-throughput CRISPR/Cas9 sgRNA validation, protein subcellular localization, and signaling pathway studies.
For rice genotypes that are recalcitrant to Agrobacterium infection, such as specific Indica varieties, we employ biolistic delivery. This physical method uses high-velocity gold or tungsten particles coated with DNA to penetrate the cell wall, delivering genetic material directly into the nucleus or chloroplasts. It is a robust alternative that bypasses biological host-pathogen compatibility barriers.
| Category | Requirements |
| Sample Type | Mature seeds, embryogenic callus, or sterile plantlets of your rice cultivar |
| Sample Amount | Minimum 50–100 g of healthy, mature seeds (approximately 2,000–3,000 seeds) |
| Pre-Treatment | Seeds should be clean, free from fungal contamination, and not chemically treated; provide detailed cultivar information |
| Storage Conditions | Store seeds at 4 °C in dry conditions; avoid prolonged storage (>6 months) |
| Shipping | Ship at ambient temperature with proper moisture control; include desiccant packets |
| Metadata Needed | Cultivar name, subspecies (indica/japonica), generation/purity, known transformation recalcitrance, target gene/construct details, preferred selection markers |
| Vector Information | Complete plasmid construct map, including promoter, gene of interest, selection marker, and reporter genes |
Complement your core transformation projects with our specialized downstream validation and precision engineering solutions to ensure high-quality research outcomes:
Molecular Characterization & Transgene Validation:
We provide comprehensive analysis to confirm successful integration and expression, including Southern Blotting for copy number determination and RT-qPCR for transcript level quantification.
CRISPR/Cas9 Off-Target Screening
To ensure the high precision of genome editing, we utilize advanced NGS-based sequencing to identify and analyze potential off-target effects across the entire plant genome.
Custom Vector Design & Construction
Our team specializes in engineering complex T-DNA vectors, including multi-gene stacking, tissue-specific promoters, and codon optimization tailored for specific host plant species.
Subcellular Localization & Imaging
We help visualize your target proteins using fluorescent tagging (GFP/YFP/RFP) and high-resolution confocal microscopy to determine precise protein distribution within plant cells.
Phenotypic Stress Tolerance Assays
Evaluate the functional impact of your genetic modifications through controlled screening for resistance to abiotic stresses like drought and salinity or biotic challenges from pathogens.
Strategy & Vector Construction
Explant Induction
Transformation & Selection
Regeneration & Hardening
Molecular Characterization
Seed Harvest
Note: Timelines may vary depending on the genotype and the complexity of the genetic modification.
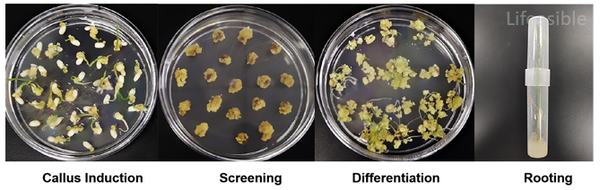
Rice Transformation for Sweet Protein Production
Internal project report highlighting the successful genetic transformation of rice (Oryza sativa) aimed at expressing sweet proteins, including Thaumatin and Brazzein. The transformation utilized Agrobacterium-mediated techniques, ensuring efficient integration of target genes into the rice genome.
The method included rigorous screening and identification protocols, confirming the presence of the desired traits in the regenerated plants.
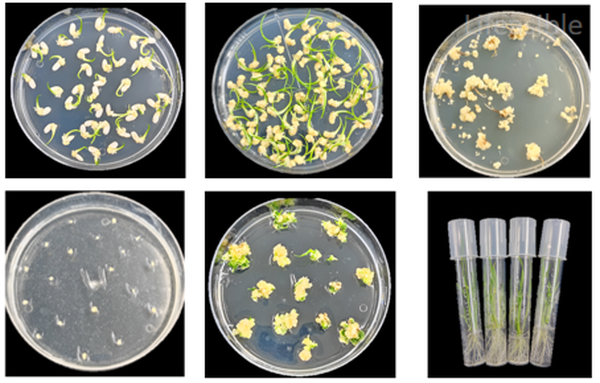

Rice Transformation using CRISPR/Cas9 Technology
This internal project report outlines the successful implementation of CRISPR/Cas9 technology for the targeted knockout of the LOC_Osxxxxxx gene in rice (Oryza sativa). The process involved designing specific sgRNAs, constructing a CRISPR vector, and transforming this vector into rice. The transformed rice callus underwent several phases of selection and regeneration.
Positive transgenic lines were verified through PCR screening, achieving a 100% success rate for the identification of positive seedlings. Subsequent editing type analysis confirmed the intended genetic modifications.
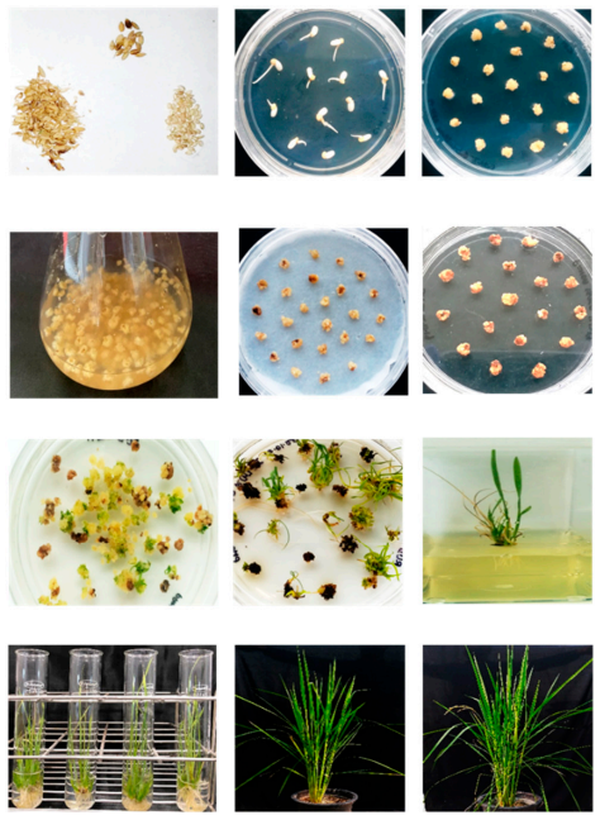

Efficient Agrobacterium-Mediated Rice Transformation for Functional Genomics
This protocol demonstrates significant optimization of Agrobacterium-mediated transformation in the japonica rice cultivar Taipei-309. By utilizing 10-12-day-old embryogenic calli rather than conventional 3-4-week-old cultures, and employing a Whatman filter paper co-cultivation system, the authors achieved transformation timelines of >60 days from seed to soil-ready plantlets under accelerated greenhouse conditions. The protocol employs N6-based media with optimized 2,4-D/BAP ratios and maltose as the primary carbon source, yielding 56-58% regeneration efficiency and 68% positive transformation rates for overexpression constructs.
This methodology was validated through two parallel applications: (i) CRISPR/Cas9-mediated knockout of OsLip1, a lipase gene implicated in rice bran rancidity, achieving 70% positive transformation efficiency with confirmed frameshift mutations; and (ii) stable overexpression of OsGolS2 (galactinol synthase), demonstrating enhanced salt-stress tolerance during seed germination. Molecular confirmation via PCR genotyping, Sanger sequencing of edited loci, and semi-quantitative RT-PCR provides a robust framework for high-throughput functional validation in rice improvement research.
Our commitment to precision and reliability has made Lifeasible a partner for academic and industrial researchers worldwide. Below are representative feedback from recent collaborations:
"Lifeasible's Agrobacterium-mediated protocol for our Indica lines yielded 12 independent T0 events, with 7 confirmed single-copy integrations via Southern blot. While we did observe some somaclonal variation in two lines, the overall efficiency allowed us to advance our drought-tolerance phenotyping ahead of schedule. Technical team was responsive to our specific vector requirements."
Dr. A. Sterling
Associate Professor
USA
"The PEG-protoplast system delivered usable data in 48 hours for 8 of our 10 sgRNA targets. Two constructs showed inconsistent results—likely due to promoter compatibility issues—but the remaining candidates proceeded to stable transformation with confirmed editing efficiency. Turnaround time was essential for our grant deadline."
Dr. L. Hoffmann
Group Leader, Plant Genome Engineering
UK
"We've commissioned 6 rice transformation projects with Lifeasible since 2021. Southern blot characterization is consistently thorough, though we recommend budgeting extra time for T2 homozygous line selection—our experience averaged 14 months from vector submission to fixed lines, slightly longer than the standard timeline. Documentation quality meets journal submission standards."
Dr. J. R. Bennett
Senior Scientist
USA
"Our elite Indica variety failed standard Agrobacterium protocols at two other service providers. Lifeasible developed a modified biolistic approach over three months, ultimately generating 6 positive T0 lines. The extended R&D phase required additional cost discussion, but transparency in troubleshooting was appreciated. Final lines are now in field trials."
Dr. M. Gauthier
Research Director
France
"For routine Nipponbare CRISPR knockouts, Lifeasible offers competitive pricing and reliable PCR genotyping. We typically receive 15-20 T0 plants per construct, with editing efficiency around 60% in our experience—lower than the advertised 80% in some cases, but sufficient for our screening needs. Recommended for labs without in-house tissue culture facilities."
Dr. R. Sinclair
Assistant Professor
USA
Rice-Specific Expertise
Decades of specialized experience in Oryza sativa transformation, ensuring deep technical knowledge of both Japonica and Indica subspecies..
Genotype Versatility
Proven success in transforming a wide range of rice varieties, from standard lab models to recalcitrant elite commercial cultivars.
Technical Precision
Industry-leading editing efficiency utilizing the latest Prime Editing (PE) and CRISPR/Cas9 technologies tailored for the rice genome.
Global Compliance
All rice engineering projects are conducted in state-of-the-art facilities that adhere to the strictest international biosafety regulations.
Are you ready to accelerate your rice research?
Our technical experts are available to discuss your project requirements, from vector design to greenhouse management. From CRISPR-based gene editing to stable transgenic line development, Lifeasible is your trusted partner for every stage of rice genetic engineering.
Oryza sativa (rice) is not only a primary global food staple but also the premier model organism for monocot research.
Rice transformation technologies have evolved significantly from early low-efficiency methods to highly optimized, genotype-flexible systems. Initial approaches relied heavily on protoplast transformation and particle bombardment, which often resulted in unstable integrations. The introduction of Agrobacterium-mediated transformation marked a major breakthrough, enabling more precise and stable gene transfer. More recently, advances such as CRISPR/Cas-based genome editing and in planta transformation strategies have further expanded capabilities, allowing faster, more accurate, and transgene-free modifications in rice research and breeding programs.
Agrobacterium-mediated transformation is widely regarded as the gold standard for rice genetic engineering due to its precision, efficiency, and stability. This method exploits the natural ability of Agrobacterium tumefaciens to transfer T-DNA into the plant genome, enabling targeted gene insertion with typically low copy numbers. Compared to physical methods, it reduces the risk of complex DNA rearrangements and gene silencing. In rice, optimized infection conditions, strain selection, and co-cultivation protocols have further improved transformation efficiency, making it highly reliable for both functional genomics and trait development.
Japonica varieties like 'Nipponbare' and 'Kitaake' are highly efficient and serve as our standard models. However, we have also optimized protocols for many Indica varieties such as '9311'.
We offer comprehensive detection of transgenic plant services, including quantitative PCR and Southern blot, to ensure data integrity.
Our standard workflow delivers T0 plantlets in 4-5 months and T1 seeds in 6-8 months from vector receipt, depending on genotype and season. For japonica model varieties like Nipponbare or Taipei-309, accelerated protocols can generate soil-ready T0 plants in 8-10 weeks under optimized greenhouse conditions. Indica varieties and complex editing projects may require 1-2 additional months due to lower regeneration efficiency.
Yes, though with modified expectations. Elite Indica varieties often exhibit poor callus induction and Agrobacterium susceptibility variability. We recommend pilot feasibility studies (10-callus batch) before full-scale commitment, with transparent reporting of genotype-specific limitations.

Extraction and Purification of Rice Genomic DNA

Detection of Genetic Diversity of Rice Parents by RFLP

Rice Anther Culture and Plant Regeneration

Understanding GMOs: A Comprehensive Guide to Genetic Modification in Agriculture
Reference